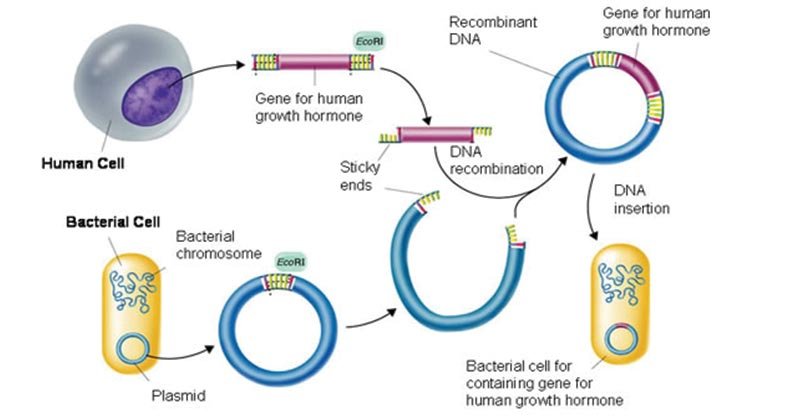
Introduction to cloning and gene analysis
Cloning gene is a common practice in molecular biology laboratories used by researchers to create copies of a particular gene for downstream applications, such as sequencing, mutagenesis, genotyping, or heterologous expression of a protein. The traditional technique of gene cloning involves the transfer of a DNA fragment of interest from an organism to a self-replicating genetic element, such as a bacterial plasmid. This technique is commonly used today to isolate long or unstudied genes and protein expression.
A more recent technique is the use of polymerase chain reaction (PCR) to amplify a gene of interest. The advantage of using PCR over traditional gene cloning, as described above, is the decreased time needed to generate a pure sample of the gene of interest. However, gene isolation by PCR can only amplify genes with predetermined sequences. For this reason, many unstudied genes require initial gene cloning and sequencing before PCR can be performed for further analysis.
DNA sequence
DNA sequencing is often the first step in understanding the genetic makeup of an organism, helping to:
- Locate regulatory and gene sequences.
- Compare homologous genes between species
- Identify mutations
Sequencing uses biochemical methods to determine the order of nucleotide bases (adenine, guanine, cytosine, and thymine) in a DNA oligonucleotide. Knowing the sequence of a particular gene will help in further analysis to understand the function of the gene. PCR is used to amplify the gene of interest before sequencing can be done. Many biotech companies offer sequencing instruments, however, these instruments can be expensive. As a result, many researchers often perform the PCR in-house and then send their samples to sequencing labs.
Site-directed mutagenesis
Site-directed mutagenesis is a widely used procedure for the study of protein structure and function by modifying the coding DNA. Using this method, mutations can be created at any specific site in a gene whose wild-type sequence is already known. Many techniques are available to perform site-directed mutagenesis. A classic method of introducing mutations, whether single base pairs or larger insertions, deletions, or substitutions, into a DNA sequence is the Kunkel method.
The first step in any site-directed mutagenesis method is to clone the gene of interest. For Kunkel’s method, the cloned plasmid is then transformed into an Escherichia coli dut ung mutant. This strain of E. coli lacks dUTPase and uracil deglycosidase, which will ensure that the plasmid containing the gene of interest is converted to DNA that lacks Ts and contains Us instead.
The next step is to design a primer that contains the region of the gene you want to mutate, along with the mutation you want to introduce. PCR with the mutated primers can then be used to create hybrid plasmids; each plasmid will now contain one strand without the mutation and the uracil bases, and another strand with the mutation and no uracil.
The final step is to isolate this hybrid plasmid and transform it into a different strain containing the uracil-DNA glycosylase (ung) gene. Uracil deglycosidase will destroy the uracil-containing chains, leaving only the chains with its mutation. When the bacteria replicate, the resulting plasmids will contain the mutation on both strands.
Genotyped
Genotyping is the process of determining the specific DNA sequence of an individual’s genotype. This process can be achieved by various techniques such as High-Resolution Fusion Analysis (HRM) or any other mutation detection technique. All of these techniques will provide insight into the genotype of the individual, which can help determine specific sequences that can be manipulated and cloned for further analysis.
Heterologous protein expression
Heterologous protein expression uses gene cloning to express a protein of interest on a self-replicating genetic element, such as a bacterial plasmid. Heterologous expression is used to produce large quantities of a protein of interest for functional and biochemical analyses.
